 |
 |
 |
 |
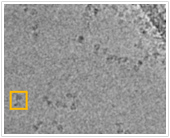


The experimental technique of Cryo Electron Microscopy (Cryo-EM) complements those of X-Ray Crystallography and Nuclear Magnetic Resonance Spectroscopy for the experimental determination of protein structure. Cryo-EM is best known for its ability to determine the structures of extremely large proteins and protein complexes (i.e., the ribosome); however, structures determined by Cryo-EM are typically of lower resolution than those determined by X-Ray Crystallography or NMR. The image at left is not white-noise, it is imaging data from a Single Particle Cryo-EM experiment. The dark shape within the orange box corresponds to a two-dimensional projection of a single protein. By observing a large number of two-dimensional projections we can reconstruct the protein's structure. Unfortunately, the relative orientations of the protein for each projection are unknown - One of the most difficult tasks in Cryo-EM is the estimation of these image parameters. Working in close collaboration with Professor John Rubinstein, we are developing algorithms to estimate these imaging parameters and to reconstruct protein structure.
For the Non-Biologist: Structural Biologists use knowledge of protein structure and function to gain insights into the molecular basis of disease and to design new pharmacologic interventions. One method for the determination of protein structure is Cryo-EM. A simple analogy demonstrates the intuition behind the method. Imagine sending a very tiny photographer to take thousands of pictures of the target protein. Upon completion, the diligent photographer presents you with several thousand photographs each annotated with the angle from which it was taken. It is not unreasonable to expect that you could construct a pretty accurate model of the protein's structure. This is the core idea behind Single Particle Cryo-EM. Unfortunately two problems lead to significant complications. First, the angle annotation of each image is lost. Second, the photographs are extremely noisy (i.e., extremely grainy). Consequently, we are left with thousands of noisy images each with an unknown imaging angle. This is what makes Cryo-EM difficult. In our research, we are developing algorithms to automatically estimate the viewing angles and then to assemble these noisy images into a three-dimensional model of protein structure.
Students: Navdeep Jaitly, Marcus Brubaker, Zongyi Yang
Collaborators: John Rubinstein (Dept of Biochemistry & Hospital for Sick Children)
|
 |
 |
 |
 |
 |
 |
 |
 |
Our goal is to develop techniques capable of obtaining better than 8A resolution structures for non-symmetric proteins. From the perspective of information content, there is sufficient information to achieve this goal with existing experimental techniques. In theory, a sufficiently large set of single-particle Cryo-EM images would contain sufficient information for a 2-3A resolution structure.  In practice, this goal has not yet been achieved. Typically, non-symmetric Cryo-EM structures are determined to 20-30A resolution. Proteins with high degrees of internal symmetry allow for more accurate estimation of projection pose. These proteins can be reconstructed at higher resolution and have been reported at beyond 8A resolution. In practice, this goal has not yet been achieved. Typically, non-symmetric Cryo-EM structures are determined to 20-30A resolution. Proteins with high degrees of internal symmetry allow for more accurate estimation of projection pose. These proteins can be reconstructed at higher resolution and have been reported at beyond 8A resolution.
The primary challenge is that of pose estimation. Post estimation is complicated by the  extremely noisy images and the effect of the microscope's Contrast Transfer Function (CTF). Combining ideas from Machine Vision, Machine Learning, and Biochemistry we are developing novel algorithms for pose estimation and model building. Using a Bayesian approach, we are looking to identify the model that best explains the data. With the long range goal of completely automating the process of Cryo-EM data analysis, we are tackling two different challenges: extremely noisy images and the effect of the microscope's Contrast Transfer Function (CTF). Combining ideas from Machine Vision, Machine Learning, and Biochemistry we are developing novel algorithms for pose estimation and model building. Using a Bayesian approach, we are looking to identify the model that best explains the data. With the long range goal of completely automating the process of Cryo-EM data analysis, we are tackling two different challenges:
 Generating an Initial Low-Resolution Model: Current techniques for model refinement require prior information on protein structure in the form of a low-resolution structural model. Existing methods for generating this low-resolution model have a significant manual component. We are interested in developing a turn-key solution, where an initial model can be constructed from a set of unprocessed particle images. Generating an Initial Low-Resolution Model: Current techniques for model refinement require prior information on protein structure in the form of a low-resolution structural model. Existing methods for generating this low-resolution model have a significant manual component. We are interested in developing a turn-key solution, where an initial model can be constructed from a set of unprocessed particle images. Model Refinement and Generation of a High-Resolution Structure: A high-resolution structure can be created from dozens of particle images if their relative orientations are known. The typical approach is iterative where estimation of projection pose and incremental model building alternate. We are casting this process into a Bayesian framework. Model Refinement and Generation of a High-Resolution Structure: A high-resolution structure can be created from dozens of particle images if their relative orientations are known. The typical approach is iterative where estimation of projection pose and incremental model building alternate. We are casting this process into a Bayesian framework.
UPDATE
In December 2009, we completed version 0.1 of our work.
 A manuscript has been submitted for review A manuscript has been submitted for review We are preparing a freely available software distribution We are preparing a freely available software distribution We are completing a parallel implementation of the software We are completing a parallel implementation of the software
Please check back here and at http://compbio.cs.toronto.edu/cryoem.

|
 |
 |
 |
 |
|
|
|