Computational Biology
SCARPA: Scaffolding Reads with Practical Algorithms
Abstract:
Current assemblers for high throughput sequencing platforms can produce
quality draft assemblies for small genomes. In contrast, the assemblies
produced for complex genomes using short reads are typically very fragmented.
The finishing stage, which requires additional sequencing, can significantly
benefit from scaffolding. We have developed Scarpa, which combines
fixed-parameter tractable and bounded algorithms with Linear
Programming in order to produce accurate scaffolds. Scarpa also
estimates library size distribution and detects mis-assembled contigs.
Publication:
Donmez N., Brudno M. "SCARPA: scaffolding reads with practical algorithms";
Bioinformatics. 2013 Feb 15;29(4):428-34. doi: 10.1093/bioinformatics/bts716.
[pdf]

Genome assembly for highly polymorphic genomes
Abstract:
Several recently sequenced genomes exhibit very high polymorphism rates.
For these organisms, genome assembly remains a challenge. Single Nucleotide
Polymorphisms (SNPs) and small indels (insertion/deletion) may be mistaken
for sequencing errors and larger variations such as Copy Number Variations
(CNVs) or rearrangements make it difficult to assemble a single reference
sequence. We introduce Hapsembler, an assembler to facilitate the assembly
of such genomes. Hapsembler features a haplotype-aware error correction
procedure and uses a novel structure called a mate pair graph to resolve
ambiguities that arise from polymorphism and repeats.
Publication:
Donmez N., Brudno M. "Hapsembler: An assembler for highly polymorphic genomes";
International Conference of Research in Computational Biology (RECOMB) 2011,
V. Bafna and S.C. Sahinalp (Eds.), LNBI 6577:38-52, 2011. [pdf]

Polymorphism in Ciona savignyi

Abstract:
We compare two haploid genotypes of one C. savignyi individual and
identify codons at which these genotypes differ by two non-synonymous
substitutions. Using the C. intestinalis genome as an outgroup, we show
that both substitutions tend to occur in the same genotype. Only in 53
(34.4%) of 154 codons, one substitution occurs in each of the two genotypes,
although 77 (50.0%) of such codons are to be expected if substitutions were
independent. We consider two feasible evolutionary causes for the observed
pattern: substitutions driven by positive selection and compensatory
substitutions, as well as several potential biases.
Publication:
Donmez N., Bazykin G., Brudno M., Kondrashov A.S. "Polymorphism due
to multiple amino acid substitutions at a codon site within Ciona savignyi";
Genetics 181:685-690, 2009. [pdf]
Graphics
Concepture: recognizing gestures with repetitive patterns
Abstract:
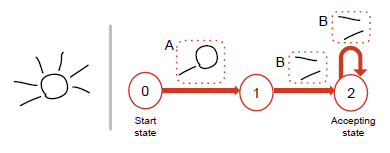
We present Concepture, a framework based on regular language grammars for the authoring and recognition
of sketched gestures with infinitely varying and repetitive patterns. Such gestures, while often seen in gesture
based applications are currently hard-coded and not customizable. We endorse an example-based workflow, where
users author gestures by sketching one or more example instances of the gesture. We algorithmically deconstruct
these examples into perceptible stroke segments. Adjacent segment-pairs further capture local spatial relationships
between segments and these segment-pairs form the alphabet of a regular language. We then initialize a grammar
for our gesture by admitting strings that represent the user provided examples. Grammar refinement is userfriendly,
in that we automatically generate new candidate gestures that are visually presented to the user for
verification as instances of the gesture. We show Concepture to be effective in efficiently authoring a number of
common, yet difficult to recognize gestures, and illustrate it using clip-art and image annotation applications.
Publication:
Donmez N., Singh K. "Concepture: a regular language based framework for
recognizing gestures with repetitive and variational patterns";
Proceedings of SBIM '12 Proceedings of the International Symposium on Sketch-Based Interfaces and Modeling.
(Best Paper Award) [pdf]